Putting the spotlight on translation initiation
A new method by the Valen lab reveals modes and regulation of translation initiation

Hovedinnhold
Most of the cell’s energy is expended for protein synthesis. Despite this, it has been a major challenge to monitor protein synthesis on individual RNAs across the whole transcriptome. This changed with the introduction of ribosome profiling, a method that captured all translating ribosomes and mapped them back to their exact position in the transcriptome. Since its inception this technique has become a standard for studying translational regulation. Despite its usefulness, ribosome profiling has a major limitation: it is completely blind to what happens during the initiation phase of translation. Initiation is a rate limiting step in protein synthesis and it is here, during this phase, that much of the process is regulated. This drawback therefore results in a blindspot for researchers trying to understand how protein synthesis is controlled.
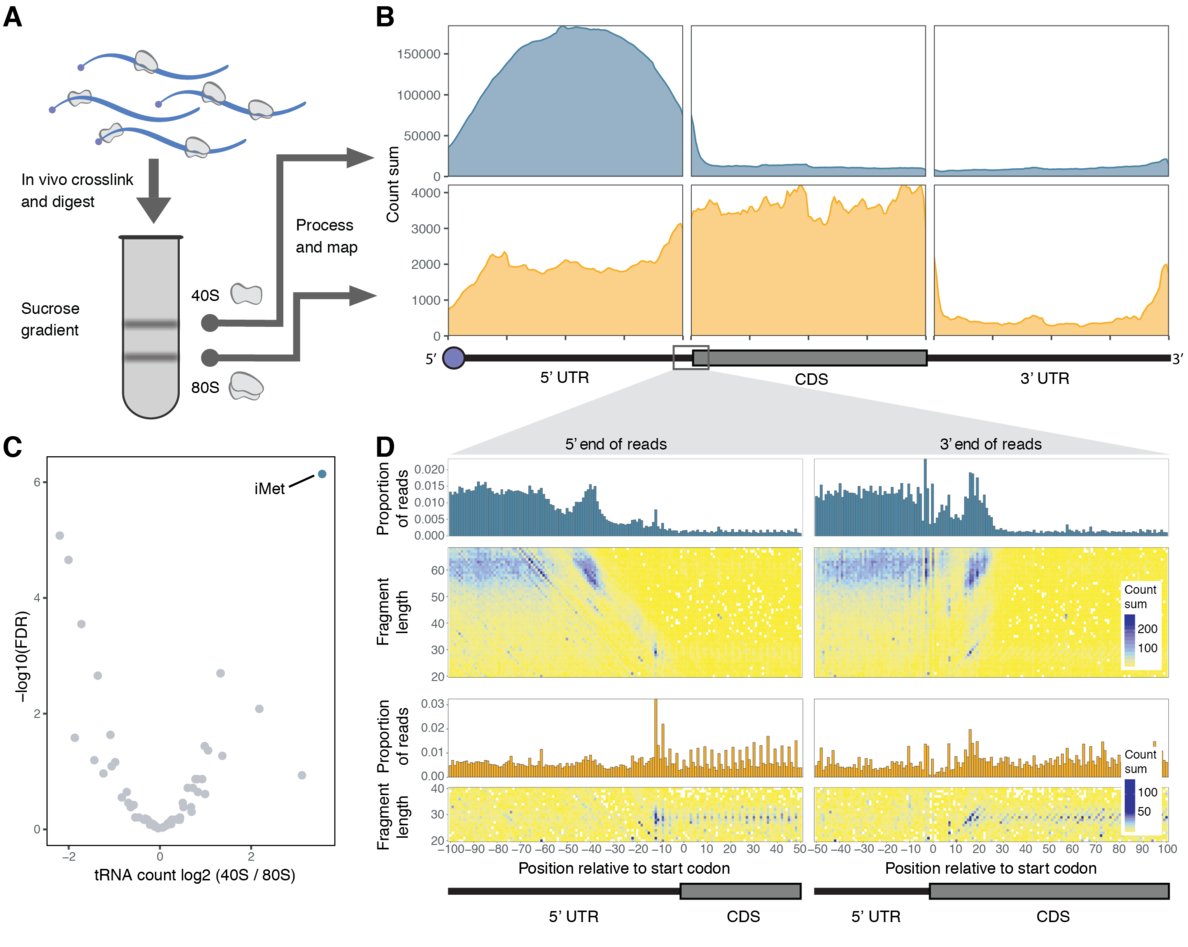
A new study from the Valen Lab puts the spotlight onto translation initiation. Published in Cell Reports, this work introduces a new method called Ribosome Complex Profiling (RCP-seq) to study scanning ribosomes as they initiate translation. Spanning both computational and experimental work, the study is the product of two Valen lab members: Adam Giess and Yamila Torres Cleuren. The study sheds light on how ribosomes are recruited to RNAs, and quantifies how effectively they traverse and scan through the 5’ UTRs. Like ribosome profiling, RCP-seq also captures all translating ribosomes and can therefore measure how scanning ribosomes recognize the start codon and progress to the elongation phase. This new perspective revealed how cells can modulate initiation by changing the context around the start codon. Together, RCP-seq presents a new window into translation initiation and is poised to reveal many of the secrets of how initiation is regulated.
Shared first author of this publication, Dr. Yamila Torres Cleuren tells about her work:
-Can you tell us about your educational and scientific background and what interested you about working with the Valen lab?
I have been lucky to study and work in several institutions and countries. Getting my MSc at King’s College London, I learned about many different fields of research in molecular biology and my curiosity for genetics really started. I did my PhD between New Zealand and California in C. elegans early development, using natural variation and population genetics to understand regulatory networks. During this time, I relied a lot on bioinformatic and statistical analyses and I got to learn a whole new set of skills. I visited the Sars Centre a few times during my PhD, as my husband was based in Bergen, and I fell in love with the city. I met with Eivind Valen (my current boss) and was so intrigued by what he was doing that I asked to join his group for my postdoc. It was the perfect opportunity to continue working with both wet lab and bioinformatics methods and learn a new field: RNA biology and translation.
-What are your main interests in science, and how do you see your current work correlating with these interests?
What intrigues me to this date is how whole organisms are formed from what are basically “blobs” of cells. Not just that, but how they reproducibly make organisms with varying external conditions. When we think of genes, we immediately think of their ultimate function - making proteins. However, this process is controlled by a wide range of players and their involvement can be fine-tuned to respond to external stimuli (e.g., infection) or internal stimuli (e.g., stress). For example, under normal conditions you might have gene A being made into mRNA and this is efficiently made into protein. But, under stressful conditions, the mRNA of gene A is not translated efficiently and instead it is repressed and it could even be that the mRNA gets degraded. Understanding how this process of translation regulation occurs and under what conditions is one of the main questions of my current research.
A problem we often encounter is that what we study is limited by the methods available to us. This is why when we hit a wall with our knowledge, we need to spend time developing new ways to continue. In the last few years, in the Valen lab we have worked very hard to come up with new methods (or variations of available methods) to study new features of RNA and translation. Our main focus is on how translation starts (initiation step) and what regulates it. Finding new ways to do research is an incredibly challenging task but a very fulfilling way to do science. Not only do you get to develop a new method, but you also get to be the first one to do this experiment and make this first observation. It’s extremely rewarding to get to do this, bring this new knowledge back to the organism and try to further understand it.
- Why would you say this new paper is important in your current research field and what should be done after it?
By being able to study all stages of translation, we were able to shed light on several questions that had been controversial. First, prior to our study, two competing models existed for the binding of ribosomes to mRNA: “threading” and “slotting”. Our data revealed that the majority of mRNAs in vivo recruit small subunits through the threading model. Second, we were able to estimate how often endogenous scanning ribosomes manage to traverse the whole 5’UTR and reach the start codon, finding that it is on average very high. We were also able to reveal features of mRNAs that impede this progress. Third, the roles of upstream Open Reading Frames (uORFs) is still heavily debated and how much impact they have on regulating translation. By providing a new way to measure this, we were able to show that uORFs are one of the major features that impede scanning ribosomes. Finally, by contrasting how much scanning we observed relative to translation, we were able to build a model for how initiation contexts impact translation. A surprise to us was that the optimal context was not the well-known Kozak sequence, and that many good contexts differed substantially from this.
Although we found new features of translational regulation and introduced new ways of analyzing these effects with RCP-seq, we have yet to explore its full potential. We want to use RCP-seq to really understand how translation is regulated under different circumstances, for example in stress and disease. The RCP-seq protocol can be further applied to study samples with limited input material, which will allow addressing heterogeneity and specialization of the translation machinery, compartmentalized translation or tissue-specific translation. This opens for the possibility to obtain novel insights into scanning and initiating mechanisms across organisms and disease models.
-What are your main interests outside science?
I love to be outdoors, especially hiking and to do water sports. I’m not the biggest fan of cold water so I only really go into the water in the summer months here, but luckily Bergen is surrounded by some amazing mountains and there’s endless tracks to explore. I love the outdoors culture in Norway. It truly is a hiker’s paradise. What I am most amazed by is how you find people of all ages out in nature, any time of the year, with all types of weather.