Computational chemistry services
Main content
As part of Nor-Openscreen and EU-Openscreen, we offer the following computational chemistry services:
HTS analysis - get more value from your screen
Based on the hit list and the compound structures in the screening set, we can analyze your screen to give you information about structure-activity relationships (SAR). This can help you to assess the value of the hit compounds and to guide their further evaluation.
Hit expansion - explore SAR
We can screen for analogues of your hit compounds in databases with commercially available compounds. This will help you to quickly explore SAR and to improve the affinities of your hits.
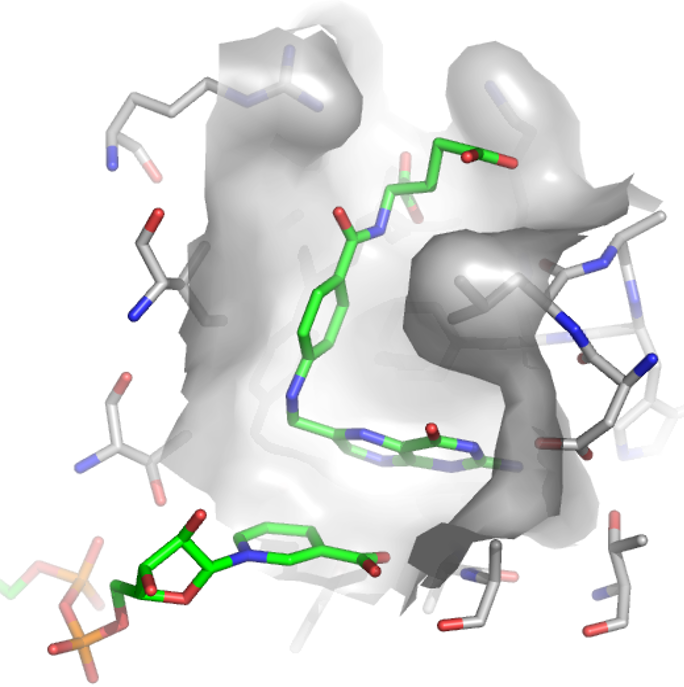
Virtual screening - find new ligands
In virtual screening, computer stored compound databases or screened for compounds that are either similar to a query compound or that fit into the binding site of a given target. The goal is the generate a list of compounds which are enriched in ligands.
Bespoke services
We also offer other bespoke services, e.g. druggability assessment, homology modeling, and compound design for hit optimization.
For all of the services, we have access to an in-house curated library of about 7 million commercially available compounds, to other publicly accessible compound database, in house developed software as well as commercial software packages.
For further information and to discuss your needs, contact Ruth Brenk.