News archive for Martinez lab
An important European-funded initiative has been launched to explore how common molecular mechanisms may link metabolic disorders, especially type 2 diabetes and obesity, with brain disorders such as Alzheimer's disease, obsessive-compulsive disorder, and autism spectrum disorders. Jan Haavik and Aurora Martinez from the Department of Biomedicine are the norwegian participants.
The Workshop in Applied Structural Bioinformatics will be organized at the University of Bergen October 16-20. This workshop will provide a practical introduction to popular tools for biomolecular visualisation, analysis and simulations. Lectures will be given by both international and local speakers from the research fields of structural bioinformatics and computer-aided drug design.
Sarsia Seed recently announced their new seed funds in Bergen consisting of 300 million NOK. Initial investment can be in the biotechnology company Pluvia founded by Prof. Aurora Martinez according to Sarsia officials.
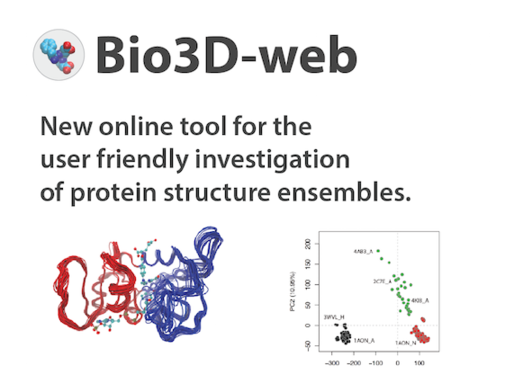
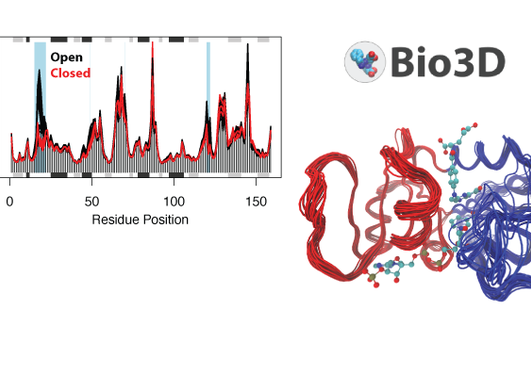
Bio3D-web is a new online tool for the analyses of sequence, structure and conformational heterogeneity of protein families. The tool is developed by researcher Dr. Lars Skjærven, member of the the Biorecognition group, in collaboration with Prof. Barry J. Grant at the University of Michigan.
The Bio3D package includes new methods for the analysis and visualization of protein dynamics both from experimental and simulated data, integrated with tools for comparative analysis of evolutionary related protein structures and systematic retrieval of publicly available sequence and structural data.
The Biorecognition group at the Department of Biomedicine represents one of the four nodes in the new NOR-Openscreen infrastructure. The infrastructure will support the discovery of biologically active substances in all areas of the Life Sciences by providing transnational, open access to the most advanced technologies, chemical and biological resources as well as expertise through Europe.
- February 2020 (1)
- January 2020 (1)
- October 2017 (1)
- March 2017 (1)
- November 2016 (1)
- August 2016 (1)
- June 2016 (1)
- June 2015 (1)